Network Pharmacology and Molecular Docking Study of Yupingfeng Powder in the Treatment of Allergic Diseases.
Study Design
- Loại nghiên cứu
- Other
- Đối tượng nghiên cứu
- None
- Can thiệp
- Network Pharmacology and Molecular Docking Study of Yupingfeng Powder in the Treatment of Allergic Diseases. None
- Đối chứng
- None
- Kết quả chính
- None
- Xu hướng hiệu quả
- Mixed
- Nguy cơ sai lệch
- Unclear
Abstract
OBJECTIVE: To explore the potential mechanisms of Yupingfeng Powder (YPFP) in the treatment of allergic diseases by using network pharmacology and molecular docking technology. METHODS: The active components and targets of YPFP were screened by the TCMSP database. The targets associated with atopic dermatitis, asthma, allergic rhinitis, and food allergy were obtained from GeneCards and OMIM databases, respectively. The intersection of the above disease-related targets was identified as allergy-related targets. Then, allergy-related targets and YPFP-related targets were crossed to obtain the potential targets of YPFP for allergy treatment. A protein-protein-interaction (PPI) network and a drug-target-disease topology network were constructed to screen hub targets and key ingredients. Next, GO and KEGG pathway enrichment analyses were performed separately on the potential targets and hub targets to identify the biological processes and signaling pathways involved. Finally, molecular docking was conducted to verify the binding affinity between key ingredients and hub targets. RESULTS: In this study, 45 active ingredients were identified from YPFP, and 48 allergy-related targets were predicted by network pharmacology. IL6, TNF, IL1B, PTGS2, CXCL8, JUN, CCL2, IL10, IFNG, and IL4 were screened as hub targets by the PPI network. However, quercetin, kaempferol, wogonin, formononetin, and 7-O-methylisomucronulatol were identified as key ingredients by the drug-target-disease topological network. GO and KEGG pathway enrichment analysis indicated that the therapeutic effect of YPFP on allergy involved multiple biological processes and signaling pathways, including positive regulation of fever generation, positive regulation of neuroinflammatory response, vascular endothelial growth factor production, negative regulation of cytokine production involved in immune response, positive regulation of mononuclear cell migration, type 2 immune response, and negative regulation of lipid storage. Molecular docking verified that all the key ingredients had good binding affinity with hub targets. CONCLUSION: This study revealed the key ingredients, hub targets, and potential mechanisms of YPFP antiallergy, and these data can provide some theoretical basis for subsequent allergy treatment and drug development.
Tóm lược
This study revealed the key ingredients, hub targets, and potential mechanisms of YPFP antiallergy, and these data can provide some theoretical basis for subsequent allergy treatment and drug development.
Full Text
Research Article Network Pharmacology and Molecular Docking Study of Yupingfeng Powder in the Treatment of Allergic Diseases
Minye Qu ,1 Wenhua Tao ,1 and Jian Ma 2
- 1Department of Traditional Chinese Medicine, First Clinical Medical College, Jiangsu University, Zhenjiang, China
- 2Department of Warm Disease, Basic Medical College, Nanjing University of Chinese Medicine, Nanjing, China Correspondence should be addressed to Minye Qu; [email protected] Received 28 March 2022; Accepted 22 June 2022; Published 9 July 2022 Academic Editor: Shih-Chao Lin
Copyright © 2022 Minye Qu et al. is is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Objective. To explore the potential mechanisms of Yupingfeng Powder (YPFP) in the treatment of allergic diseases by using network pharmacology and molecular docking technology. Methods. e active components and targets of YPFP were screened by the TCMSP database. e targets associated with atopic dermatitis, asthma, allergic rhinitis, and food allergy were obtained from GeneCards and OMIM databases, respectively. e intersection of the above disease-related targets was identified as allergyrelated targets. en, allergy-related targets and YPFP-related targets were crossed to obtain the potential targets of YPFP for allergy treatment. A protein-protein-interaction (PPI) network and a drug-target-disease topology network were constructed to screen hub targets and key ingredients. Next, GO and KEGG pathway enrichment analyses were performed separately on the potential targets and hub targets to identify the biological processes and signaling pathways involved. Finally, molecular docking was conducted to verify the binding affinity between key ingredients and hub targets. Results. In this study, 45 active ingredients were identified from YPFP, and 48 allergy-related targets were predicted by network pharmacology. IL6, TNF, IL1B, PTGS2, CXCL8, JUN, CCL2, IL10, IFNG, and IL4 were screened as hub targets by the PPI network. However, quercetin, kaempferol, wogonin, formononetin, and 7-O-methylisomucronulatol were identified as key ingredients by the drug-target-disease topological network. GO and KEGG pathway enrichment analysis indicated that the therapeutic effect of YPFP on allergy involved multiple biological processes and signaling pathways, including positive regulation of fever generation, positive regulation of neuroinflammatory response, vascular endothelial growth factor production, negative regulation of cytokine production involved in immune response, positive regulation of mononuclear cell migration, type 2 immune response, and negative regulation of lipid storage. Molecular docking verified that all the key ingredients had good binding affinity with hub targets. Conclusion. is study revealed the key ingredients, hub targets, and potential mechanisms of YPFP antiallergy, and these data can provide some theoretical basis for subsequent allergy treatment and drug development.
1. Introduction
Allergies are a class of recurrent, chronic inflammatory diseases. Over the past few decades, the incidence of allergic diseases has continued to rise, especially in developing countries [1]. Many cross-sectional and longitudinal studies have shown that the occurrence of allergic diseases follows a natural process, including the time evolution from preschool atopic dermatitis and food allergy to school-age asthma and allergic rhinitis, as well as the spatial evolution from the skin and gastrointestinal tract to respiratory tract [2, 3]. With the consensus of the natural process of allergy, the whole-course
management of allergy has been advocated in recent years [4]. However, currently available therapeutic approaches against allergy, such as glucocorticoids, antihistamines, leukotriene inhibitors, and oral immunotherapy can only temporarily control symptoms, but cannot effectively control disease recurrence or inhibit the atopic progression [5, 6]. In addition, long-term use of these drugs has certain side effects [6–8].
Traditional Chinese medicine (TCM) has a long history of treating allergic diseases. e “holistic concept” of TCM emphasizes the correlation between the various components of the human body, which coincides with allergy whole-
course management concept of modern medicine. Due to the good clinical efficacy and few side effects, TCM treatment of allergy is widely used in China and has been gradually accepted worldwide as a major treatment for complementary and alternative medicine [9, 10].
Yupingfeng Powder (YPFP) originated from the ancient Chinese medicine book “Danxi Xinfa,” which is composed of three Chinese herbs: Huangqi, Fangfeng, and Baizhu. In allergic diseases, YPFP shows good effects on both improving symptoms and controlling recurrence [11–13]. It has been confirmed that the glycoside extract of YPFP exerts antiinflammatory and immunological functions by regulating T lymphocyte differentiation subtypes [14]. e Huangqi-Fangfeng drug pair prevents allergic airway remodeling by inhibiting the epithelial-mesenchymal transformation process via regulating the epithelial-derived TGF-β1 [15]. e polysaccharide isolated from Baizhu enhances immunity by promoting the proliferation of CD4, CD8, and Treg cells [16, 17]. However, the TCM compound has the characteristics of multiple components, multiple targets, and multiple pathways. e pharmacodynamic components, effective targets, and molecular mechanisms of YPFP on allergy remain to be further studied.
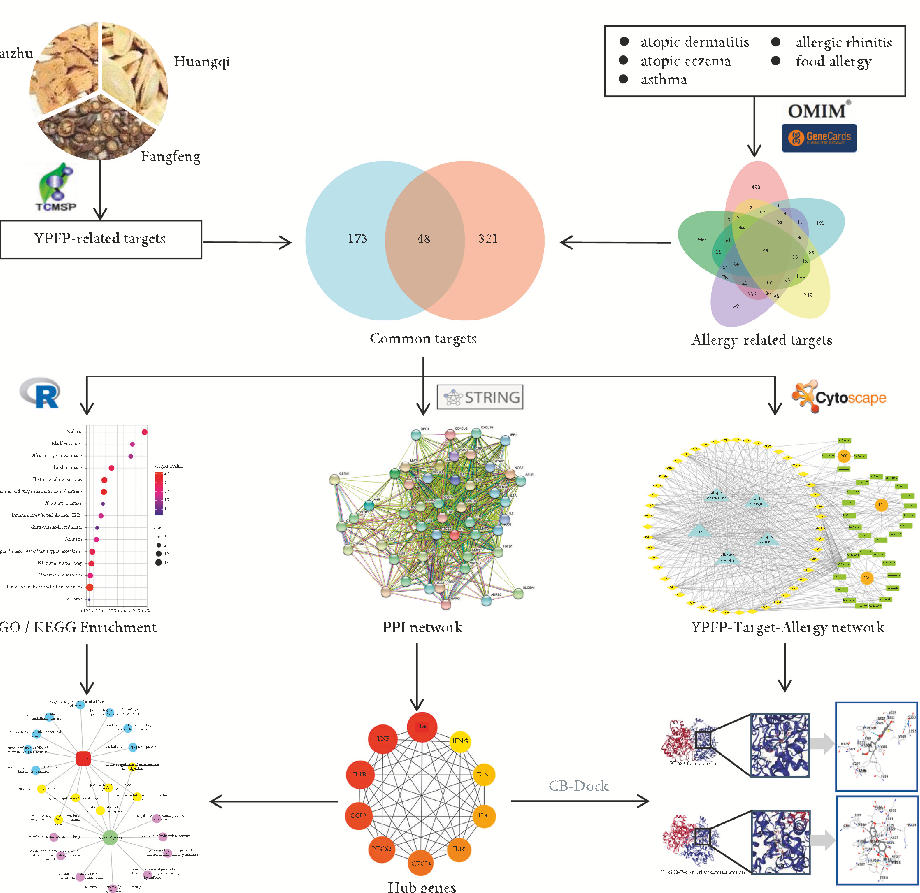
With the rapid development of system biology and system pharmacology, network pharmacology has transformed the drug research models from “single disease-single target” to “multiple disease-multiple target”, which provides a new way for us to systematically study the pharmacodynamic mechanism of TCM compounds [18, 19]. erefore, in this study, network pharmacology was conducted to predict the active ingredients of YPFP as well as its targets and pathways for allergy treatment. In addition, molecular docking techniques were used to verify the interactions between ingredients and targets. is study aimed to provide a theoretical basis for the potential mechanism of YPFP antiallergy. e workflow is shown in Figure 1.
2. Methods
- 2.1. Active Ingredient Screening of YPFP. e active ingredients of YPFP were obtained from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP) (https://tcmspw.com/tcmsp.php), which is a unique TCM systematic pharmacology platform that integrates TCM active ingredients, potential targets, associated diseases, and pharmacokinetic data. Using “Huangqi,” “Fangfeng,” and “Baizhu” as keywords, oral bioavailability (OB) ≥30% and drug-likeness (DL) ≥0.18 as screening conditions, a total of 45 active ingredients were obtained. Download the molecular structures of the screened active ingredients to prepare for subsequent molecular docking.
- 2.2. Target Prediction of Active Ingredients. Targets corresponding to the active ingredients of YPFP were predicted from TCMSP and translated into gene names using UniProtKB (https://www.uniprot.org). After removing duplicate genes, 221 YPFP-related target genes were obtained.
- 2.3. Allergy-Related Target Gene Collection. Allergy-related target genes were searched for the keywords “atopic dermatitis,” “atopic eczema,” “asthma,” “allergic rhinitis,” and “food allergy” by GeneCards (https://www. genecards.org) and OMIM (https://www.omim.org). e data from the two databases were merged to remove the duplicates, and the related genes for each keyword were obtained. Cross the five keyword-related genes by “UpSet” and “VennDiagram” R package to obtain allergy-related target genes.
- 2.4. Clustering of YPFP and Allergy-Related Target Genes. Venny2.1 (https://bioinfogp.cnb.csic.es/tools/venny) was used to cluster YPFP-related target genes and allergy-related target genes, and the obtained overlaps were considered as potential target genes of YPFP therapy for allergy.
- 2.5. PPI Network Construction and Hub Gene Extraction. e PPI network was plotted by entering the potential target genes into the STRING database (https://www.string-db. org), with the species set as “Homo sapiens” and a confidence score ≥0.4. Each node in the network represents a protein, and the connection lines between nodes represent functional associations. e relevant results were exported in TSV format and extracted by Cytoscape v_3.9.1, and the top 10 hub genes were identified by the McCreight (MCC) method using the “Cytohubba” plug-in.
- 2.6. Construction of YPFP-Target-Allergy Topological Network. Cytoscape v_3.9.1 was used to analyze and visualize complex networks among herbs, active ingredients, potential targets, and diseases. By analyzing the network topology parameters, the key ingredients were screened.
- 2.7. GO and KEGG Pathway Enrichment Analysis. KOBAS 3.0 (https://KOBAS.cbi.pku.edu.cn) was used for the KEGG and GO analyses of the potential genes and hub genes, respectively. e enrichment degree was calculated according to the input number/background number, with P value ≤0.05 and an input number ≥2 as the inclusion criteria. Based on descending order of enrichment degree, the top 15 KEGG pathway and top 15 GO items of potential genes and hub genes were separately plotted as bubble diagrams by the “ggplot2” R package. en Cytoscape v_3.9.1 was used to construct topological networks to identify the common biological functions and signaling pathways of hub genes and potential genes in the top 15 GO and KEGG results. In addition, the “ggalluvial” R package was used to draw the Sankey diagram of hub genes and enriched signal pathways. KEGG Mapper (https://www.genome.jp/kegg/ mapper) is a collection of tools for KEGG mapping, which visualizes the relevant pathways of acquired genes through “Search Pathway” and “Search & Color Pathway” tools. We input potential genes into KEGG Mapper and set different colors (the hub genes are red, the remaining potential genes are green), and then got the most relevant signaling pathway and displayed them.
atopic dermatitis atopic eczema asthma
allergic rhinitis food allergy
Baizhu
Huangqi
Fangfeng
YPFP-related targets
Common targets Allergy-related targets
GO / KEGG Enrichment PPI network YPFP-Target-Allergy network
CB-Dock
Hub genes
pathways and functions
Figure 1: Workflow of this study.
- 2.8. Molecular Docking of Hub Targets and Key Ingredients. To verify the binding affinity of the hub targets of the PPI network to the key ingredients of the topological network, we performed molecular docking. CB-Dock is an online molecular docking website (https://clab.labshare.cn/cbdock/php/). It predicts the binding region of a given protein, calculates the center and size using a curvation-based cavity detection method, connects with ligand binding sites queried by Autodock Vina, and then sorts binding patterns based on Vina scores and provides interactive 3D visualization of binding patterns. e 3D structures of the hub targets obtained by RSCB PDB and the 2D structures of the key ingredients obtained by TCMSP were uploaded to the CB-Dock website. After automatically identifying cavity size,
calculating center and size, molecular docking, and configurational scoring were performed. Cavity size is the protein-ligand interaction interface predicted by CB-Dock based on the concave surface of protein [20]. Generally, the larger the cavity size is, the higher the affinity between protein and ligand will be [20]. e center and size of the docking box are evaluated according to the cavity center, cavity size, and ligand size, which are key parameters in the process of molecular docking [21]. e binding affinity was evaluated according to the Vina score of the docking results. e greater the absolute value of the Vina score, the more stable the docking module is. e 3D conformation of the target-ingredient docking with a Vina score≤−8.5kcal/mol was downloaded for display.
Table 1: Characteristics of active ingredients in YPFP. Herb name Molecule ID Molecule name
Molecule weight
OB (%)
DL
MOL000211 Mairin 456.78 55.38 0.78 MOL000239 Jaranol 314.31 50.83 0.29 MOL000296 Hederagenin 414.79 36.91 0.75
(3S, 8S, 9S, 10R, 13R, 14S, 17R)-10, 13-Dimethyl-17-[(2R, 5S)-5-propan-2yloctan-2-yl]-2, 3, 4, 7, 8, 9, 11, 12, 14, 15, 16, 17-dodecahydro-1H-cyclopenta[a] phenanthren-3-ol
MOL000033
428.82 36.23 0.78
MOL000354 Isorhamnetin 316.28 49.6 0.31 MOL000371 3, 9-di-O-methylnissolin 314.36 53.74 0.48 MOL000374 5′-Hydroxyiso-muronulatol-2′, 5′-di-O-glucoside 642.67 41.72 0.69
- MOL000378 7-O-Methylisomucronulatol 316.38 74.69 0.3
- MOL000379 9,10-Dimethoxypterocarpan-3-O-β-D-glucoside 462.49 36.74 0.92
- MOL000380
Huangqi (HQ)
(6aR, 11aR)-9, 10-Dimethoxy-6a, 11a-dihydro-6H-benzofurano[3, 2-c] chromen-3-ol
300.33 64.26 0.42
MOL000387 Bifendate 418.38 31.1 0.67 MOL000392 Formononetin 268.28 69.67 0.21 MOL000398 Isoflavanone 316.33 109.99 0.3 MOL000417 Calycosin 284.28 47.75 0.24 MOL000422 Kaempferol 286.25 41.88 0.24 MOL000433 FA 441.45 68.96 0.71
- MOL000438 (3R)-3-(2-Hydroxy-3,4-dimethoxyphenyl)chroman-7-ol 302.35 67.67 0.26
- MOL000439 Isomucronulatol-7, 2′-di-O-glucosiole 626.67 49.28 0.62 MOL000442 1, 7-Dihydroxy-3, 9-dimethoxy pterocarpene 314.31 39.05 0.48 MOL000098 Quercetin 302.25 46.43 0.28
(2R, 3R)-3-(4-Hydroxy-3-methoxy-phenyl)-5-methoxy-2-methylol-2, 3dihydropyrano[5, 6-h][1,4] benzodioxin-9-one
386.38 68.83 0.66
MOL000011
MOL011730 11-Hydroxy-sec-o-beta-d-glucosylhamaudol_qt 292.31 50.24 0.27 MOL011732 Anomalin 426.5 59.65 0.66 MOL011737 Divaricatacid 320.32 87 0.32 MOL011740 Divaricatol 334.35 31.65 0.38
- MOL001941 Ammidin 270.3 34.55 0.22 MOL011747 Ledebouriellol 374.42 32.05 0.51 MOL011749 Phelloptorin 300.33 43.39 0.28 MOL011753 5-O-Methylvisamminol 290.34 37.99 0.25 MOL002644 Phellopterin 300.33 40.19 0.28 MOL000359 Sitosterol 414.79 36.91 0.75 MOL000173 Wogonin 284.28 30.68 0.23 MOL000358 Beta-sitosterol 414.79 36.91 0.75 MOL001494 Mandenol 308.56 42 0.19
- MOL001942 Isoimperatorin 270.3 45.46 0.23 MOL003588 Prangenidin 270.3 36.31 0.22 MOL007514 Methyl icosa-11,14-dienoate 322.59 39.67 0.23 MOL013077 Decursin 328.39 39.27 0.38
Fangfeng (FF)
- MOL000020 12-Senecioyl-2E, 8E, 10E-atractylentriol 312.39 62.4 0.22
- MOL000021 14-Acetyl-12-senecioyl-2E, 8E, 10E-atractylentriol 355.44 60.31 0.31
- MOL000022 14-Acetyl-12-senecioyl-2E, 8Z, 10E-atractylentriol 356.45 63.37 0.3 MOL000028 α-Amyrin 426.8 39.51 0.76
Baizhu (BZ)
(3S, 8S, 9S, 10R, 13R, 14S, 17R)-10, 13-Dimethyl-17-[(2R, 5S)-5-propan-2yloctan-2-yl]-2, 3, 4, 7, 8, 9, 11, 12, 14, 15, 16, 17-dodecahydro-1H-cyclopenta[a] phenanthren-3-ol
MOL000033
428.82 36.23 0.78
MOL000049 3β-Acetoxyatractylone 274.39 54.07 0.22 MOL000072 8β-Ethoxy atractylenolide III 276.41 35.95 0.21
3. Results
- 3.1. Active Ingredients and Targets Screening of YPFP. According to OB≥30% and DL≥0.18, a total of 45 active ingredients of YPFP were screened from the TCMSP database, including 20 from Huangqi, 18 from Fangfeng, and 7 from Baizhu (Table 1). By TCMSP target prediction and
UniProt gene name transformation, 221 YPFP-related target genes were obtained after removing duplicates (Supplementary Table S1).
3.2. Allergy-Related Target Gene Collection. e GeneCards and OMIM databases were searched with the keywords
800
759
452
600
150
12
47
40 20
12
17
10 4
192
62
460452
460
51
369
35
28
400
369
3318
345
59
22
326
78 67 122
23 17
326 20
65 345
759
192
200
150
122
78 67 65 62 59 51 47 40 35 33 28 23 22 20 20 18 17 17 12 12 10 4 0
atopic dermatitis
food allergy
asthma
atopic eczema
allergic rhinitis
2000 1500 1000 500 0 Set Size
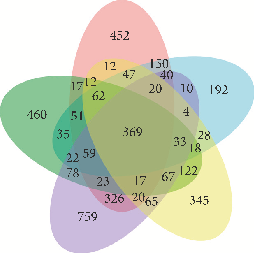
- Figure 2: Allergy-related target genes obtained by clustering target genes corresponding to “atopic dermatitis,” “atopic eczema,” “asthma,” “allergic rhinitis,” and “food allergy.”
“atopic dermatitis,” “atopic eczema,” “asthma,” “allergic rhinitis,” and “food allergy.” When the search results of the two databases were combined (if relevance score >1, duplicates were removed), the target genes corresponding to the 5 keywords were 1140, 1677, 1445, 1912, and 1241, respectively (Supplementary Table S2). By clustering the genes associated with the 5 keywords, 369 shared genes were obtained as allergy-related target genes (Figure 2).
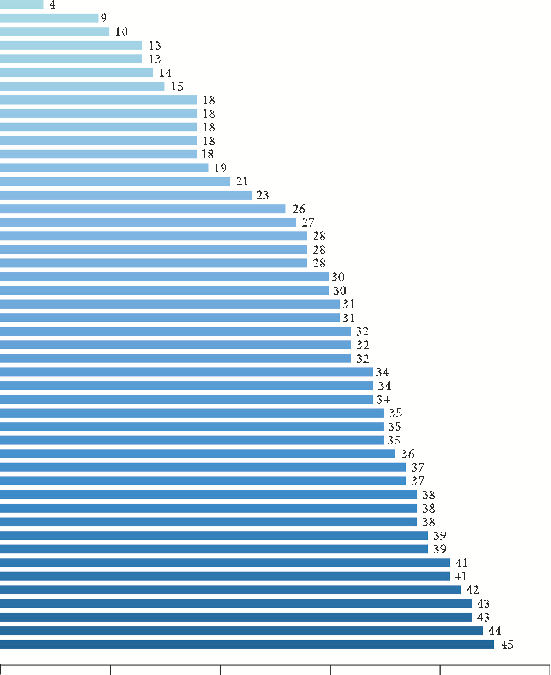
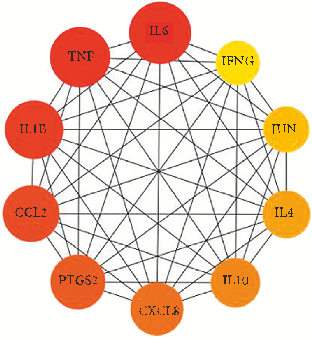
- 3.3. Potential Target Genes and PPI Network of YPFP erapy for Allergy. YPFP-related target genes and allergy-related target genes were intersected by Venny2.1 to obtain 48 potential genes (Figure 3(a)). ese 48 potential genes were imported into the STRING database to obtain the PPI network (Figure 3(b)). According to the node degree of each protein in the network, the importance priority was analyzed and a bar graph was drawn for visualization (Figure 3(c), Supplementary Table S3). In order to screen hub genes from the potential genes, PPI network files were imported into Cytoscape v_3.9.1, and the top 10 genes were identified as hub genes by the MCC method of the “CytoHubba” plug-in, including IL6, TNF, IL1B, PTGS2, CXCL8, JUN, CCL2, IL10, IFNG, and IL4 (Figure 3(d), Supplementary Table S4).
- 3.4. YPFP-Target-Allergy Topological Network. Cytoscape v_3.9.1 was used to construct a topological network to
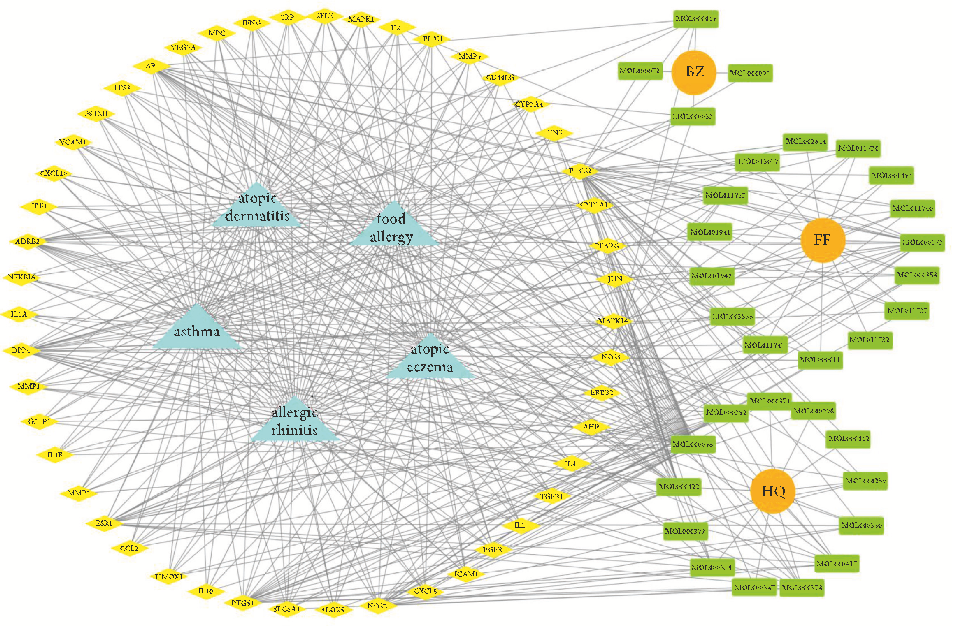
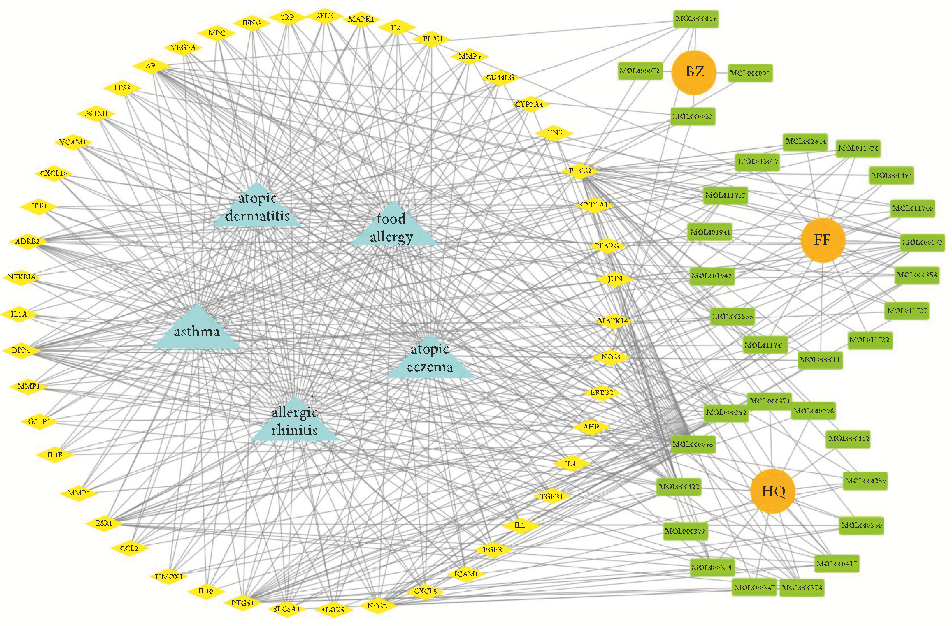
visualize the complex relationship between YPFP active ingredients and allergy-related targets. e YPFP-TargetAllergy network consists of 88 nodes and 478 interactions. ese nodes were 32 YPFP active ingredients, 48 potential targets, 5 keywords of allergy-related diseases, and 3 herbs (Figure 4). e active ingredients were sorted according to the descending order of degree value, and the top 5 were predicted to be the key ingredients, including quercetin, kaempferol, wogonin, formononetin, and 7-O-methylisomucronulatol (Table 2).
3.5. GO and KEGG Pathway Enrichment Analysis of Potential Genes and Hub Genes. To explore the regulatory mechanisms of potential target genes on allergies, we conducted GO and KEGG enrichment analyses and selected the top 15 remarkably enriched biological functions and signal pathways to generate bubble diagrams, respectively (Figure 5).
To further verify the dominant role of hub genes in potential genes, we also performed GO and KEGG enrichment analyses on hub genes and compared the results with those of potential genes. e GO and KEGG enrichment results of the top 15 hub genes were represented by bubble diagrams (Figure 6). In addition, the common enriched pathways and functions of hub genes and potential genes in the top 15 GO and KEGG analyses were displayed by topological networks (Figure 7).
TCM Disease
173 48 321
- (a)
- (b)
Figure 3: Continued.
SLC6A4 GSTM1 ADRB2
4
- 9
- 10 13
CYP3A4 PTGS1 GSTP1
- 13
- 14
- 15 18 18 18 18
CYP1A1 ALOX5 AR
DPP4 MAPK1
PLAU IRF1 AHR
- 18
- 19 21
CD40LG MAPK14
23
- 26
- 27
- 28 28 28
ERBB2 MMP1
NOS2
SELE CXCL10
- 30
NOS3 ESR1 IL1A
CRP MPO
TGFB1 HMOX1
- 34
- 35
- 36
- 37
- 38
- 39 41
IL2 MMP2
EGFR
ICAM1 VCAM1 NFKBIA
IFNG
IL4 CCL2
IL10 MMP9
JUN PPARG CXCL8
- 41
- 42
- 43
VEGFA
PTGS2 IL1B TP53
- 43
- 44
- 45
TNF IL6
0
10 20 30 40 50
(c)
IL6 TNF
IFNG
IL1B
JUN
CCL2
IL4
PTGS2
IL10
CXCL8
- Figure 3: Potential target genes and PPI network of YPFP therapy for allergy. (a) e Venny results of potential target genes of YPFP therapy for allergy. (b) e PPI network of 48 potential genes. (c) Count and list of the genes of the PPI network. (d) Hub genes from the PPI network.
GO enrichment analysis of hub genes and potential genes showed that 7 of the top 15 biological functions with the highest enrichment were identical, such as positive regulation of fever generation, positive regulation of neuroinflammatory response, vascular endothelial growth factor production, negative regulation of cytokine production
involved in immune response, positive regulation of mononuclear cell migration, type 2 immune response, and negative regulation of lipid storage.
KEGG enrichment analysis showed that 11 of the top 15 signaling pathways with significant enrichment of hub genes and potential genes were the same, including malaria,
CRP SELE MAPK1
MOL000049
IFNG
IL6
MPO
PLAU
VEGFA
MMP9
AR
BZ
MOL000072 MOL000022
CD40LG
TP53
CYP3A4
GSTM1
MOL000033
TNF
MOL002644
VCAM1
MOL011730
MOL013077
PTGS2
CXCL10
MOL001494
atopic dermatitis
MOL011753
CYP1A1
IRF1
MOL011740
food allergy
- MOL001941
- MOL001942
FF MOL000173
ADRB2
PPARG
MOL000358
NFKBIA
JUN
MOL011737
IL1A
MOL003588
MAPK14
asthma
MOL011732
atopic eczema
MOL011747
DPP4
NOS3
MOL000011
MMP1
ERBB2
MOL000371 MOL000392 MOL000296
allergic rhinitis
GSTP1
AHR
MOL000442
MOL000098
IL1B
IL4
MOL000239
HQ
MOL000422
MMP2
TGFB1
ESR1
IL2
MOL000380
MOL000379
CCL2
EGFR
MOL000417
MOL000354
ICAM1 CXCL8
HMOX1
MOL000387 MOL000378
IL10
PTGS1
SLC6A4 ALOX5 NOS2
- Figure 4: YPFP-Target-Allergy network. e orange circles represent drugs; the green squares represent active ingredients; the yellow diamonds represent potential targets; and the blue triangles represent the diseases. e line between two nodes represents the interaction.
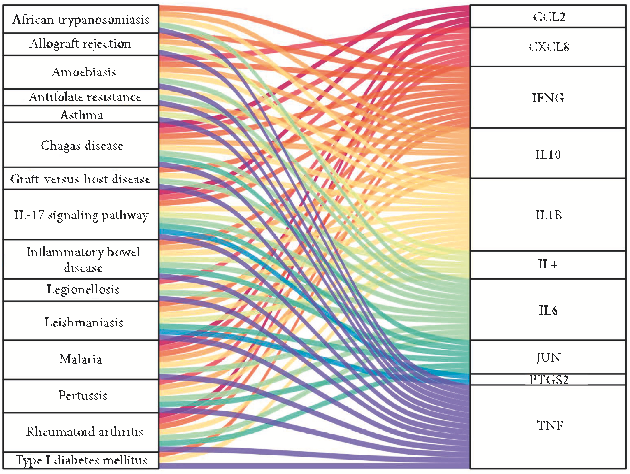
African trypanosomiasis, inflammatory bowel disease (IBD), allograft rejection, graft-versus-host disease, IL-17 signaling pathway, asthma, leishmaniasis, pertussis, chagas disease (American trypanosomiasis), and rheumatoid arthritis. Additionally, the Sankey diagram showed that most hub genes were enriched in the IL-17 pathway. e Sankey diagram and signal pathway diagram are shown in Figures 8 and 9.
Table 2: Top 5 high degree active ingredients in YPFP-TargetAllergy network.
No Molecule id Molecule name (active ingredients) Degree
- 1 MOL000098 Quercetin 41
- 2 MOL000422 Kaempferol 21
- 3 MOL000173 Wogonin 17
- 4 MOL000392 Formononetin 14
- 5 MOL000378 7-O-Methylisomucronulatol 11
- 3.6. Molecular Docking Verification. Molecular docking was used to evaluate the binding affinity between the hub targets of the PPI network and the key ingredients of the YPFPTarget-Allergy topological network. e results showed that the minimum vina score of each docking module was less than −5.0kcal/mol, indicating a good binding affinity of the hub targets to the key ingredients (Table 3, Supplementary Table S5). To better demonstrate the docking patterns of the targets and ingredients, only the docking modules with a vina score of less than −8.5kcal/mol were visualized (Figure 10).
- 4. Discussion
Atopic dermatitis (also known as atopic eczema), food allergies, asthma, and allergic rhinitis are the most common allergy-related diseases with similar epigenetic and physiopathological characteristics [22, 23]. Numerous
experimental and clinical studies have shown that the poor prognosis of atopic dermatitis and food allergies in early life greatly exacerbates the occurrence of later asthma and allergic rhinitis [1–4, 24]. Although it is believed that the development of allergies is associated with genetic and environmental influences, the specific mechanisms remain unclear. erefore, it is necessary to conduct joint research on multiple allergy-related diseases to explore their common core mechanisms.
TCM has a unique theoretical system for the treatment of allergies. According to the theory of TCM, “qi deficiency” and “wind evil” are the two pathogenic bases of allergic diseases. erefore, invigorating qi and dispelling wind are the main treatment principles. YPFP, a TCM compound with the above dual effects, is considered an effective and safe complementary and alternative therapy for allergy. To reveal the underlying mechanism of YPFP for allergy treatment, we described the relationships between active ingredients,
positive regulation of fever generation negative regulation of lipid storage
vascular endothelial growth factor production positive regulation of neuroinflammatory response
-log10(pvalue)
8 7 6 5
type 2 immune response
regulation of immunoglobulin secretion positive regulation of mononuclear cell migration
positive regulation of vascular endothelial cell proliferation negative regulation of production of miRNAs involved in gene silencing by miRNA
count
2.0 2.5 3.0 3.5 4.0
negative regulation of gene silencing by miRNA negative regulation of cytokine production involved in immune response
membrane to membrane docking estrogen 2-hydroxylase activity connective tissue replacement involved in inflammatory response wound healing regulation of cytokine production involved in inflammatory response
0.40 0.45 0.50 0.55 0.60
(a)
Malaria
Bladder cancer African trypanosomiasis
-log10(pvalue)
Leishmaniasis IL-17 signaling pathway
30 25 20 15 10
AGE-RAGE signaling pathway in diabetic complications
Allogra rejection Inflammatory bowel disease (IBD) Gra -versus-host disease Pertussis Chagas disease (American trypanosomiasis) TNF signaling pathway Rheumatoid arthritis Fluid shear stress and atherosclerosis Asthma
count 4 8 12 16
0.125 0.150 0.1750.200 0.2250.250
(b)
- Figure 5: e top 15 remarkably enriched GO and KEGG pathway analyses of potential genes. (a) e top 15 remarkably enriched GO analyses for biological function of potential target genes of YPFP in allergy. (b) e top 15 remarkably enriched KEGG analyses for signaling pathway of potential target genes of YPFP in allergy.
targets, and signaling pathways in combination with network pharmacology and molecular docking.
A PPI network was constructed with 48 genes coexpressed by allergy and YPFP. e 10 hub genes obtained by the PPI network all suggest the association of allergy and inflammatory response. IL6, TNF, CCL2, CXCL8, and IL1β
were pro-inflammatory genes, while IL4, IL10, and IFNG were antiinflammatory genes. In the acute phase of inflammation, IL-6 is an important mediator of host immunity, but its sustained release can induce chronic inflammation or even a “cytokine storm” [25]. Mast cells (MC) are key participants in IgE lazy allergy [26]. In local reactions, MC-mediated CCL2
positive regulation of fever generation
vascular endothelial growth factor production positive regulation of neuroinflammatory response
-log10(pvalue)
type 2 immune response positive regulation of mononuclear cell migration
9 8 7 6
negative regulation of cytokine production involved in immune response postive regulation of NMDA glutamate receptor activity positive regulation of MHC class II biosynthetic process
count
positive regulation of prostaglandin biosynthetic process response to molecule of bacterial origin regulation of establishment of endothelial barrier negative regulation of amyloid-beta clearance
2.00 2.25 2.50 2.75 3.00
endothelial cell apoptotic process negative regulation of lipid storage
positive regulation of membrane protein ectodomain proteolysis
0.2 0.3 0.4 0.5 0.6
(a)
Malaria
-log10(pvalue)
African trypanosomiasis
Inflammatory bowel disease (IBD)
Allograft rejection
Graft-versus-host disease
- 3
- 4
- 5
- 6
- 7
- 8
- 9
- 10
IL-17 signaling pathway
Asthma
Antifolate resistance
count
Leishmaniasis
Pertussis
Chagas disease (American trypanosomiasis)
Rheumatoid arthritis
Legionellosis
Type I diabetes mellitus
Amoebiasis
0.06 0.08 0.10 0.12 0.14
(b)
- Figure 6: e top 15 remarkably enriched GO and KEGG pathway analyses of hub genes. (a) e top 15 remarkably enriched GO analyses for biological function of hub genes of YPFP in allergy. (b) e top 15 remarkably enriched KEGG analyses for signaling pathway of hub genes of YPFP in allergy.
promotes basophil migration, and MC-mediated CXCL8 and IL-1β participate in the recruitment of neutrophils [27–29]. Besides, TNF, as an important derivative of MC, plays a key role in promoting the induction of adaptive immunity [26]. TH2 type adaptive immunity is a common immunological manifestation of allergy-related diseases [30]. TH2 polarization leads to dysregulation of T lymphocyte subsets, such as TH1/TH2 imbalance and TH2/Treg imbalance [31, 32]. As the main secreted cytokines of TH2, TH1, and Treg, the expressions of IL-4, IFN-c, and IL-10 necessarily show
corresponding changes [31, 32]. PTGS2 is a cyclooxygenase involved in pain, inflammation, and tumor formation. Recent studies have found that PTGS2 participates in the induction of neovascularization and the promotion of TH2 polarization [33–35]. Furthermore, Jun regulates inflammatory processes at the transcriptional level. It was confirmed that Jun/AP-1 activation plays a key role in the expression of pro-inflammatory molecules such as IL-1, IL-6, and TNF-A [36]. us, anaphylaxis is essentially a chronic inflammatory response involving multiple cells and mediators.
negative regulation of amyloid-beta clearance regulation of establishment of endothelial barrier
positive regulation of membrane protein ectodomain proteolysis
positive regulation of prostaglandin biosynthetic process
response to molecule of bacterial origin
endothelial cell apoptotic process
positive regulation of NMDA glutamate receptor activity
hub genes
positive regulation of mononuclear cell migration
positive regulation of MHC class II biosynthetic process
positive regulation of fever generation
type 2 immune response negative regulation of lipid storage
vascular endothelial growth factor production
positive regulation of neuroinflammatory response
regulation of immunoglobulin secretion
negative regulation of cytokine production involved in immune response
estrogen 2-hydroxylase activity
membrane to membrane docking
potential genes
regulation of cytokine production involved in inflammatory response
connectivre tissue replacement involved in inflammatory response wound healing
negative regulation of production miRNAs involved in gene silencing by miRNA
negative regulation of vascular endothelial cell proliferation
negative regulation of gene silencing by miRNA
(a)
IL-17 signaling pathway Gra -versus-host disease
Type I diabetes mellitus
Antifolate resistance
Inflammatory bowel disease (IBD)
Legionellosis
Allogra rejection
Malaria
Amoebiasis
hub genes
African trypanosomiasis
Leishmaniasis
potential genes
Bladder cancer
Pertussis
AGE-RAGE signaling pathway in diabetic complications
Chagas disease (American trypanosomiasis)
TNF signaling pathway Fluid shear stress and
Rheumatoid arthritis
Asthma atherosclerosis
(b)
- Figure 7: Co-enrichment functions and pathways of hub genes and potential genes in the top 15 GO and KEGG analyses. (a) e common enriched functions of hub genes and potential genes in the top 15 GO analyses. (b) e common enriched pathways of hub genes and potential genes in the top 15 KEGG analyses. e red square represents hub genes; the green square represents potential genes; the blue circles represent biological functions or pathways enriched by hub genes; the purple circles represent biological functions or pathways enriched by potential genes; and the yellow circles represent co-enrichment of biological functions or pathways of hub genes and potential genes.
CCL2 CXCL8
African trypanosomiasis Allogra rejection
Amoebiasis Antifolate resistance
IFNG
Asthma Chagas disease Gra -versus-host disease
IL10
IL-17 signaling pathway
IL1B
Inflammatory bowel disease Legionellosis Leishmaniasis
IL4
IL6
JUN PTGS2
Malaria
Pertussis
TNF
Rheumatoid arthritis Type I diabetes mellitus
IL-17 SIGNALING PATHWEAY
IL-17E IL-17E targeting cells producing cells
T cell, Macrophage, Nuocyte, Ih2 cell, MMP-type2 cell, Epithelial cell.
CD4+T cell, Mast cell, Eosinophil Basophil, Epithelial cell
- IL-17RA TRADD FADD
- IL-17RB Act1 TRAF6
IL-17E
IL-17A and IL-17F targeting cells CD4+T cell, CD8+T cell, yST cell, NKT cell, LTi-like cell, Epithelial cell
IL-17A and IL-17F producing cells
Epithelial cell, Keratinocyte, Endothelial cell, Macrophage, T cell, B cell, Fibroblast
+p
- IL-17A IL-17RA
- IL-17B
+u IL-17F
Act1
+p
17 cell differentiation
Cytokine-cytokine receptor interaction
USP25
IL-17C producing cells
IL-17C targeting cells
Monocyte CD4+T cell, DC,
Macrophage, Keratinocyte
IL-17B producing cells
IL-17B targeting cells
Monocyte, Endothelial cell
Condrocyte, Neuron
IL-17RB IL-17RB
Act1 TRAF6
Casp
NF-kB AP-1
Apoptosis Apoptosis
+p +p
MAPK signaling pathway
mRNA stability
+p
IL-17D producing cells
CD4+T cell, B cell
IL-17D ?
IL-17D targeting cells
Endothelial cell, Myeloid progenitor
2 cytokines DNA IL-4 IL-5 IL-13 Chemokines TARC Eotaxin
Induces 2 cell responses and suppresses 17 cell responses
DNA
Chemokines
CXCL2 CXCL5 CXCL8 CCL2 CCL7 CCL20
CXCL1 CXCL10
Cytokines
COX2
IL-6 TNFα
G-CSF GM-CSF
Anti-microbial
defensin MUC5AC MUC5B
S100A7 S100A8 S100A9 LCN2
Tissue remodeling
MMP1 MMP3 MMP9 MMP13
Autoimmune pathology Neutrophil recruitment Immunity to extracellular pathogens
DNA
DNA
Inflammatory genes
IL-6 IFN-y
TNFα
Antibacterial peptides 17 cytokines
IL-1B
Inflammatory genes
TNFα
Inflammation Host defence Autoimmune
Proinflammatory activities?
DNA
Proinflammatory activities?
04657 1/12/21 (c) Kanehisa Laboratories
5. Conclusion
In this study, we combined network pharmacology and molecular docking technology to study the molecular mechanism of YPFP in the treatment of allergy. According to the research, YPFP plays an antiallergic role mainly through inhibiting inflammatory infiltration, regulating neuroinflammation, regulating fever, inhibiting vascular endothelial growth, and improving lipid storage, involving malaria, African trypanosomiasis, inflammatory bowel disease (IBD), allograft rejection, graft-versus-host disease, IL-17 signaling pathway, asthma, leishmaniasis, pertussis, Chagas disease (American Trypanosomiasis) and Rheumatoid arthritis and other signaling pathways. Quercetin, Kaempferol, Wogonin, Formononetin, and 7-o-methylisomucronulatol are the key active ingredients of YPFP. IL6, TNF, IL1β, PTGS2, CXCL8, JUN, CCL2, IL10, IFNG, and IL4 are the main therapeutic targets. is study indicates that YPFP has the characteristics of multicomponent, multitarget, and multipathway in allergy treatment, which provides more evidence for the further application of YPFP in allergy.
Figures
Tables
Table 1
Table 2
Table 3
Table 4
Used In Evidence Reviews
Similar Papers
Molecules (Basel, Switzerland) · 2016
Quercetin and Its Anti-Allergic Immune Response.
The Journal of allergy and clinical immunology · 2009
Complementary and alternative medicine: herbs, phytochemicals and vitamins and their immunologic effects.
Allergy, asthma, and clinical immunology : official journal of the Canadian Society of Allergy and Clinical Immunology · 2020
Quercetin with the potential effect on allergic diseases.
Autoimmunity · 2023
Quercetin improves the imbalance of Th1/Th2 cells and Treg/Th17 cells to attenuate allergic rhinitis.
Primary care · 2002
Respiratory and allergic diseases: from upper respiratory tract infections to asthma.
Phytomedicine : international journal of phytotherapy and phytopharmacology · 2004